Research
Evolution of mobile genetic elements
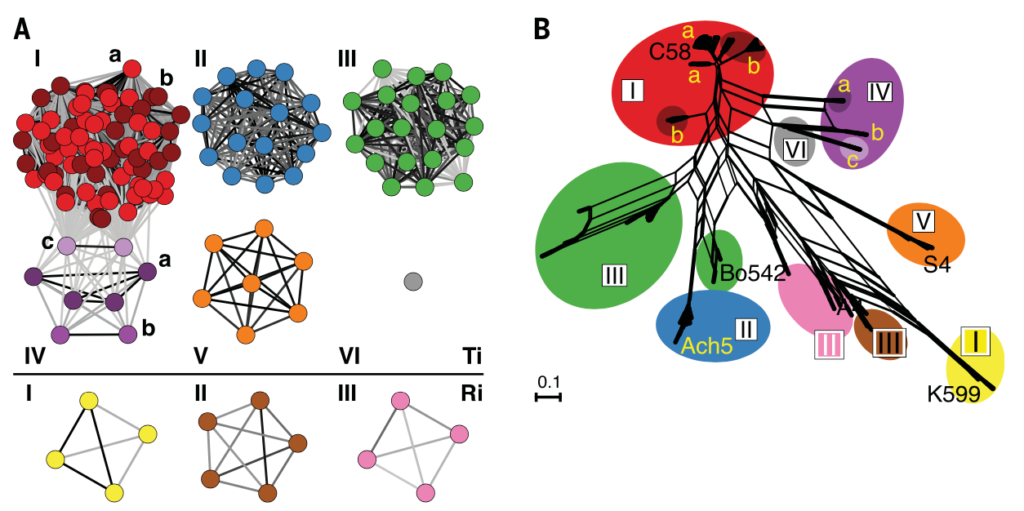
Mobile genetic elements (MGEs) play an important role in many plant/microbe interactions, as well as in human disease, such as those that carry genes conferring antibiotic resistance. MGEs, such as plasmids or integrative conjugative elements (ICEs), can cause bacterial hosts to transition from commensal organisms to pathogens or beneficial mutualists. These selfish genetic elements can recombine as well as transmit horizontally between strains, making understanding their evolution and epidemiology difficult.
What are the processes that diversify MGEs? How do they exchange loci with other elements and replicons? How do MGEs evolve and change over time?
Genomic epidemiology
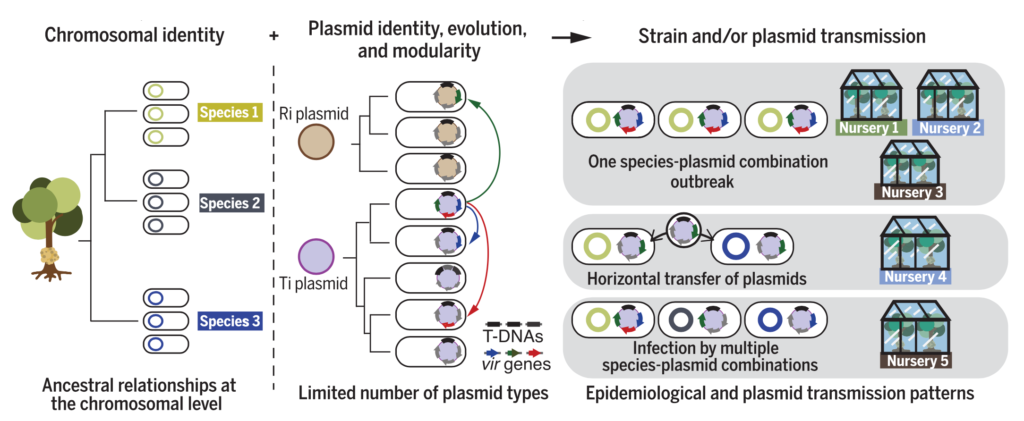
Our research investigates the genomic epidemiology of diverse agricultural pathogens, including Agrobacterium tumefaciens (crown gall disease), Rhodococcus fascians (leafy gall disease), Streptomyces scabiei (potato common scab), and Xanthomonas hortorum pv. carotae (disease of carrot), among others.
Genomic epidemiology that accurately takes MGEs into account requires generating a phylogenomic framework for accurate whole genome SNP analysis and comparison that minimizes errors and false-positive SNPs. We are developing methods to infer the epidemiology and movement of mobile genetic elements independent of their microbial hosts.
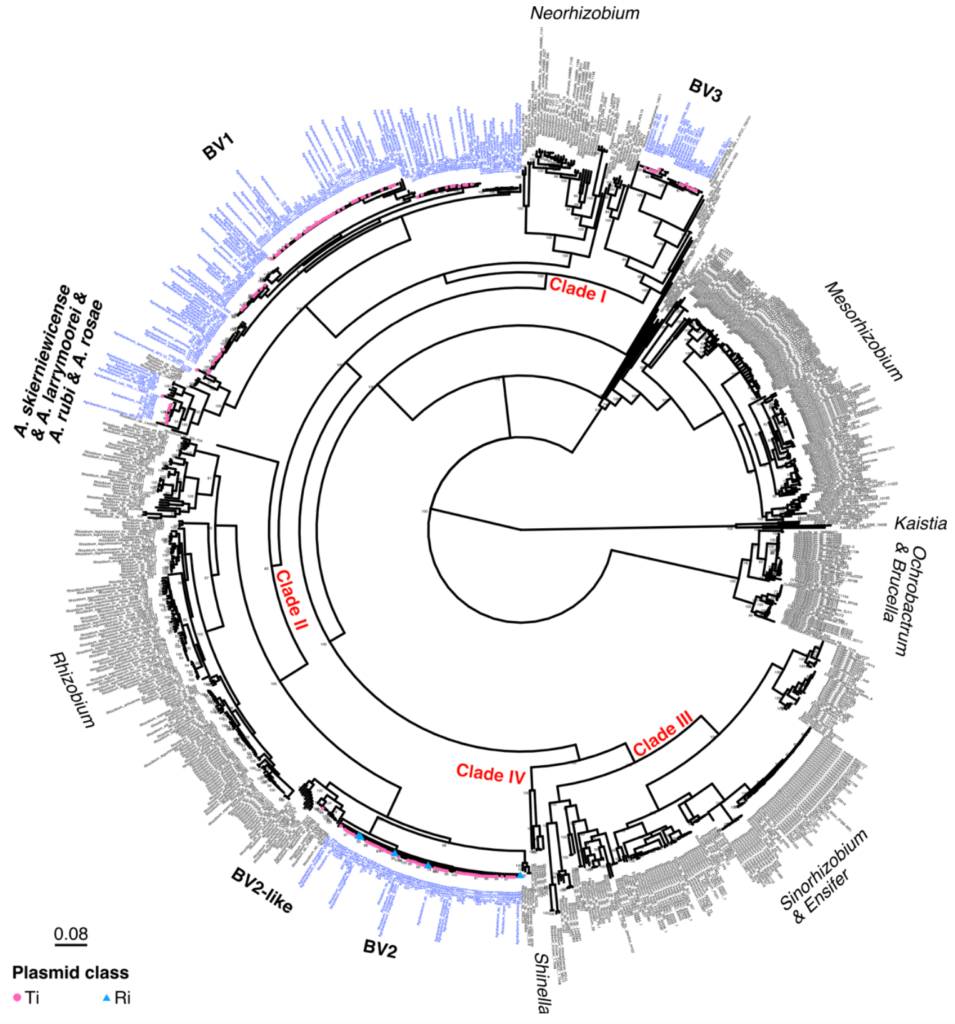
Determinants of host range and MGE compatibility
What determines the host range of a pathogen or symbiont? What confers or restricts the host range of a mobile genetic element?
Microbial pathogens (and mutualist symbionts) often have a restricted range of host species on which they can cause disease (or provide beneficial services). Some pathogens have a wide host range, while others are more restricted to one or few host species. The genes conferring this phenotype may be present on the chromosome or on mobile genetic elements, or both.
Mobile genetic elements may also be restricted to specific taxonomic groups or lineages of microbes in which they can persist.
What restricts the host range of a MGE?